Two scientists at Dartmouth College in Hanover, New Hampshire, are pioneering the first experimental use of algorithms employing structure-based molecular modeling with the objective of optimizing deimmunized drug candidates.

The work by Karl Griswold, PhD, an Associate Professor of Engineering at the Thayer School of Engineering at Dartmouth and co-investigator Christopher Bailey-Kellogg, PhD, a Professor of Computer Science at Dartmouth’s Norris Cotton Cancer Center,
Thayer School of Engineering at Dartmouth and co-investigator Christopher Bailey-Kellogg, PhD, a Professor of Computer Science at Dartmouth’s Norris Cotton Cancer Center,  complements their prior sequence-based deimmunizing algorithms and expands the tool kit of protein engineering technologies they’ve developed for deployment in next generation drug development.
complements their prior sequence-based deimmunizing algorithms and expands the tool kit of protein engineering technologies they’ve developed for deployment in next generation drug development.
Drs. Griswold and Bailey-Kellogg’s paper, “Protein Deimmunization via Structure-based Design Enables Efficient Epitope Deletion at High Mutational Loads,” (DOI: 10.1002/bit.25554) has been published in the journal Biotechnology and Bioengineering, and is coauthored by the lead corresponding authors with Regina S. Salvat of the Thayer School of Engineering, and Yoonjoo Choi of Department of Computer Science at Dartmouth College, and Alexandra Bishop of Colby College in Waterville, Maine.
The researchers note that anti-drug immune responses constitute a unique risk factor for biotherapeutics, observing that undesired immunogenicity can alter pharmacokinetics, compromise drug efficacy, and in some cases even threaten patient safety. Consequently, in order to fully capitalize on the promise of biotherapeutics, more efficient and generally applicable protein deimmunization tools will be required, and mutagenic deletion of a protein’s T cell epitopes is one powerful strategy that can be used to engineer immunotolerance. However, deimmunizing mutations must maintain protein structure and function.
In this research, EpiSweep, a structure-based protein design and deimmunization algorithm, has been used to produce a panel of seven beta-lactamase drug candidates having 27-47% reductions in predicted epitope content, which, despite bearing eight mutations each, all seven engineered enzymes maintained good stability and activity, and in the meantime variants exhibited dramatically reduced interaction with human class II major histocompatibility complex proteins, key regulators of anti-drug immune responses.
The coauthors note that when compared with 8-mutation designs generated with a sequence-based deimmunization algorithm, the structure-based designs retained greater thermostability and possessed fewer high affinity epitopes, which are dominant drivers of anti-biotherapeutic immune responses. They conclude that these experimental results validate their theory that the first structure-based deimmunization algorithm is capable of mapping optimal biotherapeutic design space, and that “by designing optimal mutations that reduce immunogenic potential while imparting favorable intramolecular interactions, broadly distributed epitopes may be simultaneously targeted using high mutational loads.”
“This work is part of our larger collaborative initiative to develop performance-enhanced protein drugs that are invisible to the human immune system,” Dr. Griswold explains in a Dartmouth Hitchcock Norris Cotton Cancer Center release. “Biotherapeutics offer potent treatment options for a wide range of diseases but, due to their biological origins, these powerful therapies can elicit detrimental immune responses in humans.”
The release notes that development of biotherapeutic agents is both time-consuming and costly, and accompanied by a substantial risk that deleterious immunogenicity issues will undermine and derail otherwise promising drug candidates late in the development process. And while methods for identifying immunogenic hotspots, or epitopes, are evolving rapidly, they observe that technologies to redesign the hotspots while maintaining biotherapeutic activity and stability are far less developed.
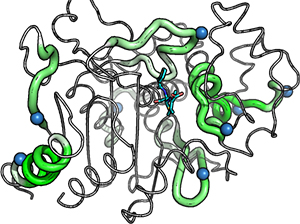
This study reported in the Biotechnology and Bioengineering paper used P99 betalactamase (the beta-lactamase of Enterobacter cloacae P99), a component of Antibody Directed Enzyme Prodrug Therapy, to demonstrate that structure-based deimmunization resulted in highly-active and stable biotherapeutic designs different from ones generated with earlier sequence-based algorithms. In particular, the structure-based designs remodeled a putative immunogenic hotspot that had proved not easily addressed using other methods.
“We demonstrated that integrating molecular modeling into deimmunizing algorithms enables simultaneous redesign of numerous immunogenic hotspots distributed throughout a protein target,” comments Dr. Bailey-Kellogg. “These results suggest that even the most immunogenic drug candidates might be engineered for improved compatibility with the human immune system.”
Dartmouth’s team of Griswold and Bailey-Kellogg have now turned their attention to advanced assays and methodologies that will allow them to better assess their deimmunized drug candidates’ immunogenic potential — methods they anticipate will yield clinically relevant data indicating to what extent they have succeeded in mitigating the immunogenicity risk of target proteins in the drug candidates. The work behind this study was funded by National Institutes of Health grant RO1-GM-098977, a Luce Foundation Fellowship, and National Science Foundation grant CNS-1205521
Sources:
Dartmouth Hitchcock Norris Cotton Cancer Center
Dartmouth College
Biotechnology and Bioengineering
Wikipedia
Image Credits:
Dartmouth Hitchcock Norris Cotton Cancer Center
Dartmouth College

